Tomato DNA Markers
The cultivated tomato (Solanum lycopersicum L.) is the second most commonly consumed vegetable crop (after potato) and unquestionably the most popular garden crop in the world. In the United States, it is the most economically important vegetable crop, and is fifth in crop value after maize, soybean, wheat, and cotton.
With the advent of molecular markers and genetic maps, there has been an increased interest in using markers technology to facilitate tomato crop improvement. Tomato was among the first crop species for which genetic markers and maps were developed and utilized for breeding purposes (Tanksley, 1983; Tanksley et al., 1992).
Tomato was one of the first crops for which molecular markers were suggested as indirect selection criteria for breeding purposes (Rick and Fobes, 1974; Tanksley and Rick, 1980; Tanksley, 1983).
More recently, with the development of new molecular markers and maps in tomato, Marker Assisted Selection- MAS has become a routine practice in many tomato breeding programs, in particular in the private sector.
Recently, several years ago, also other morphological tomato traits were mapped and now are becoming further used in MAS for genotyping and enhancing commercial breeding programs. Traits like ripening inhibitor tomato lycopene and beta carotene mutations. Plant growth structures along fruit peel and color and sugar content (BRX) traits are now routinely genotyped together with the resistance markers. Below are elaborations for several of these tomato morphological traits:

sciencedirect.com/
nor non-ripening Similar to the rin mutant, nor fruits fail to ripen and have a green pericarp [4]. The nor mutation comprises a 2 bp deletion in the third exon of NAC-NOR, resulting in a truncated protein [11]. The mutation reduces both ethylene production and lycopene biosynthesis

U Uniform ripening
Modern tomato (Solanum lycopersicum) varieties are bred for uniform ripening (u) light green fruit phenotypes to facilitate harvests of evenly ripened fruit. U encodes a Golden 2-like (GLK) transcription factor, SlGLK2, which determines chlorophyll accumulation and distribution in developing fruit.
SGR stay green, green-flesh (gf)
Fruit of the green-flesh (gf) mutant of tomato (Solanum lycopersicum) ripen to a muddy brown color due to the accumulation of lycopene coupled with a lack of chlorophyll degradation (Kerr, 1956). The lack of chlorophyll degradation in gf is not restricted to fruits because dark- and nutrient-induced chlorophyll loss in leaves is also compromised in the mutant (Akhtar et al., 1999).

Dark Green dg
Several light-hypersensitive mutants have been described in tomato (Lycopersicon esculentum). Among these, mutants carrying the monogenic recessive high pigment (hp-1 and hp-2) and dark green (dg) mutations are characterized by their exaggerated light responsiveness.
The hp-1 mutant was originally discovered as a spontaneous mutant in 1917 at the Campbell Soup Company farms (Riverton, N.J.) (Reynard 1956), the hp-2 mutant was reported in the Italian San Marzano variety in 1975 (Soressi 1975) and the dg mutant appeared in trellised planting of the Manapal variety (Konsler 1973). Despite some initial confusion, it is now clear that hp-1 and hp-2 mutations map to the tomato chromosomes 2 and 1, respectively, and are therefore non-allelic
The dg mutant is phenotypically similar to other hp mutants, but has a much darker mature-green fruit, resulting from a higher total chlorophyll content (Konsler 1973; Wann et al. 1985).

Yellow Psy1
Phytoene synthase (PSY) catalyzes the dimerization of two diterpenes from geranylgeranyl pyrophosphate to phytoene, which is the rate-limiting step in carotenoid biosynthesis. Mutations in the tomato PSY1, yellow flesh, are widely used for breeding yellow tomatoes.

Tangerine tng
Fruit of tangerine are orange and accumulate prolycopene (7Z,9Z,7′Z,9′Z-tetra-cis-lycopene) instead of the all-trans-lycopene, which normally is synthesized in the wild type.

Pink
The color of tomato fruit is mainly determined by carotenoids and flavonoids. Phenotypic analysis of an introgression line (IL) population derived from a cross between Solanum lycopersicum ‘Moneyberg’ and the wild species Solanum chmielewskii revealed three ILs with a pink fruit color. These lines had a homozygous S. chmielewskii introgression on the short arm of chromosome 1, consistent with the position of the y (yellow) mutation known to result in colorless epidermis, and hence pink-colored fruit, when combined with a red flesh. Metabolic analysis showed that pink fruit lack the ripening-dependent accumulation of the yellow-colored flavonoid naringenin chalcone in the fruit peel, while carotenoid levels are not affected.
SP self-pruning
Vegetative and reproductive phases alternate regularly during sympodial growth in tomatoes. In wild-type ‘indeterminate’ plants, inflorescences are separated by three vegetative nodes. In ‘determinate’ plants homozygous for the recessive allele of the SELF-PRUNING (SP) gene, sympodial segments develop progressively fewer nodes until the shoot is terminated by two consecutive inflorescences.
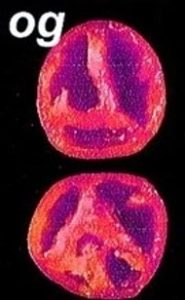
“Old-Gold” og and Beta – beta carotene
Two mutations that affect fruit pigmentation in tomato (Lycopersicon esculentum): Beta (B), a single dominant gene that increases β-carotene in the fruit, and old-gold (og), a recessive mutation that abolishes β-carotene and increases lycopene.
Curl-3 (dwarf)
Brassinosteroids (BRs) are plant steroid hormones that are essential for normal plant development. To gain better understanding of the conservation of BR signaling, the partially BR-insensitive tomato mutant altered brassinolide sensitivity1 (abs1) was identified and found to be a weak allele at the curl3 (cu3) locus. BR content is increased in both of these mutants and is associated with increased expression of Dwarf.
Brassinosteroids (BRs) are steroid hormones that affect plant growth and development, including the promotion of stem elongation, xylogenesis, and leaf bending, as reviewed by Mandava (1988) and Clouse and Sasse (1998). Mutants in the biosynthesis of the most bioactive BR, brassinolide (BL), are dark green dwarfs that are deetiolated when grown in the dark, which indicates that they lack the normal dark-grown developmental program (skotomorphogenesis). These dramatic phenotypes highlight the importance of steroid hormones in plants, and many reviews have discussed how recent advances in our understanding of BR biosynthesis and signaling are based on the analysis of such mutants.
www.researchgate.net/publication
INL internode length
Internode length is an important agronomic trait affecting plant architecture and crop yield. Plant height is an important component of plant architecture, and is highly correlated with the yield. One of the decisive factors affecting plant height is internode length. The reduced plant height or internode length of semi-dwarf varieties has improved the harvest index and biomass production.
https://pubmed.ncbi.nlm.nih.gov
LIN5 BRX
Previous studies have provided strong evidence that LIN5, a cell wall invertase, is a key determinant of the total soluble solid, or Brix, content in tomato.
Lin5 is one of a family of genes that encode invertases—enzymes that cleave sucrose to form glucose and fructose. The enzymes are classified according to their subcellular localization; invertases are present in the vacuole, cytoplasm and in the apoplast (often referred to as extracellular invertase) (Sturm and Tang 1999). Those enzymes present in the vacuole and cytosol control the sucrose : hexose ratio in the cell (Zrenner et al. 1996, Sturm 1999). The invertase in the apoplast is thought to play two main roles. First, by cleaving sucrose in the apoplast it helps to maintain a favorable sucrose gradient for unloading of sucrose from the phloem (Sturm and Tang 1999). In doing so, it increases the total capacity for uptake of sugar into the sink organ (due to the presence of both sucrose and monosaccharide transporters) (Bush 1993). Secondly, the action of apoplastic invertase is thought to be a mechanism by which sugar signals are used to integrate source and sink metabolism (Roitsch 1999). In this regard, invertase may also act to amplify sugar signals since its expression is induced by glucose, the product of its activity (Roitsch et al. 1995). The importance of apoplastic invertase in sink organ development is demonstrated by mutant and transgenic plants deficient in this enzyme that have retarded or aborted sink organ formation (Miller and Chourey 1992, Tang et al. 1999). Transgenic manipulations of invertase expression also highlight the signaling role of apoplastic invertase; expression of a heterologous invertase in potato tuber apoplast in a temporally inappropriate manner has a profound impact upon tuber morphology (Sonnewald et al. 1997).
The expression of LIN5 in the conducting tissue next to a potential “sugar-unloading site” near the ovaries is consistent with the role of LIN5 as a “sink gene” that regulates the ability of the fruit to import photosynthetic sugars.

